Fe-S Complex Enzymes
- Page ID
- 492
Callstack:
at (Workbench/LarsenLab/Research/PhotoBiology/Enzymatic_Activity/Fe-S_Complex_Enzymes), /content/body/pre, line 1, column 1
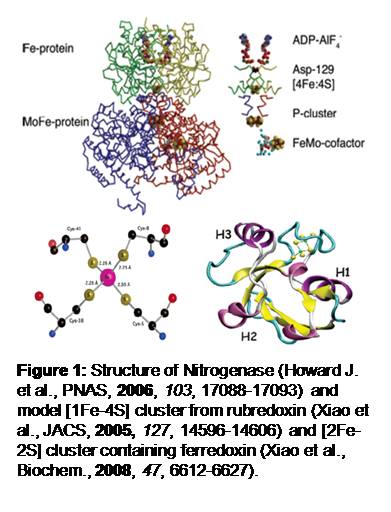 Background: In nature, enzymes utilize various transition metal clusters to catalyze chemical reactions such as electron transfer, ligand binding and its associated redox reaction, and even synthesis of other metal clusters. I am interested in Fe-S clusters found in various ferredoxin and in more complicated proteins like nitrogenase (which also has Mo containing metal cluster). Since the crucial chemical reactions happen at those metal clusters, understanding the nature of substrate binding and obtaining transient structural information during chemical reactions will be very valuable to understand how enzymes catalyze certain chemical reactions. The eventual goal of my research is to deduce transient structures of metal clusters in nitrogenase intermediates during nitrogen fixation, and elucidate mechanism of the reaction, which has been a subject to intense study for many years but yet to be determined.
Background: In nature, enzymes utilize various transition metal clusters to catalyze chemical reactions such as electron transfer, ligand binding and its associated redox reaction, and even synthesis of other metal clusters. I am interested in Fe-S clusters found in various ferredoxin and in more complicated proteins like nitrogenase (which also has Mo containing metal cluster). Since the crucial chemical reactions happen at those metal clusters, understanding the nature of substrate binding and obtaining transient structural information during chemical reactions will be very valuable to understand how enzymes catalyze certain chemical reactions. The eventual goal of my research is to deduce transient structures of metal clusters in nitrogenase intermediates during nitrogen fixation, and elucidate mechanism of the reaction, which has been a subject to intense study for many years but yet to be determined.

Techniques: In a pump probe technique, when ultrafast pulses (faster than electronically decaying time) of light are pumped on a sample, a coherent vibrational wavepacket can be generated. This coherent wavepacket contains information on vibrational modes of the system, which can be deduced by Fourier transforming the observed oscillations in the signal (in this case, the change of absorbance). This vibrational information has structural context of the system when normal mode analysis of model systems in addition to different structural data (like x-ray crystal structure and other types of vibrational spectroscopy data) construct the structural model of the system. By capturing the coherent wavepacket at different stages of the reaction, which is possible due to ultrafast light sources and advanced flow-cells, the structural model of the intermediates can be made. Below are the experimental set up and some preliminary data from one of the model system ferredoxin from Rhodobacter capsulatus.